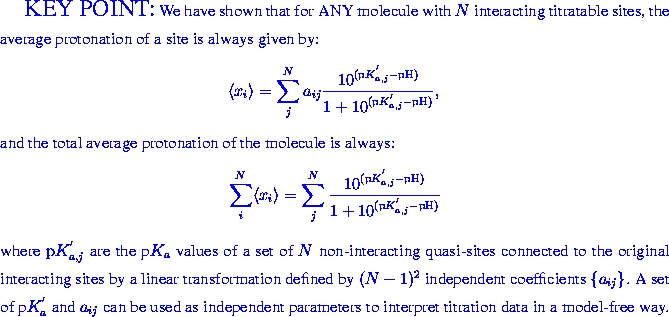
THEORY OF COOPERATIVE LIGAND BINDING
Binding of ligands to macromolecules is one of the most important reactions in biology; well-known examples include oxygen binding to hemoglobin or binding of protons to titratable groups in proteins. The binding curves may often appear deceptively featureless, yet the underlying microscopic models may be rather complex, and connections between them not intuitive.A novel view on titration of biomolecules.
When individual titratable sites in a molecule interact with each other, their pH-titration can be considerably more complex than that of an independent site described by the classical Henderson-Hasselbalch equation. We propose a novel framework - the Decoupled Sites Representation - that decomposes any complex titration behavior into simple standard components. The approach maps the set of N interacting sites in the molecule onto a set of N independent, non-interacting quasi-sites, each characterized by a pK' value. The titration curve of an individual site in the molecule is a weighted sum of Henderson-Hasselbalch curves corresponding to the quasi-sites. The total protonation curve is the unweighted sum of these Henderson-Hasselbalch curves. We show that pK' values correspond to deprotonation constants available from methods that measure total proton uptake or release, and establish their connection to protonation curves of individual residues obtained by NMR or infra-red spectroscopy. The new framework is tested on a small molecule diethylene-triamine-penta-acetate (DTPA) exhibiting non-monotonic titration curves, where it gives an excellent fit to experimental data. We demonstrate that the titration curve of a site in a group of interacting sites can be accurately reconstructed, if titration curves of the other sites are known. The application of the new framework to the protein rubredoxin demonstrates its usefulness in calculating and interpreting complicated titration curves.

APPLICATION: Titration of DTPA.
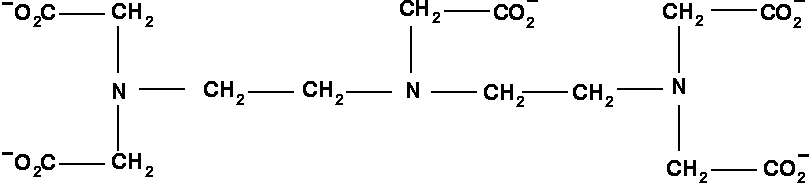
- LEFT PANEL: Structural formula of
DTPA. Only three nitrogens titrate within the range of interest from pH=3 to 13.
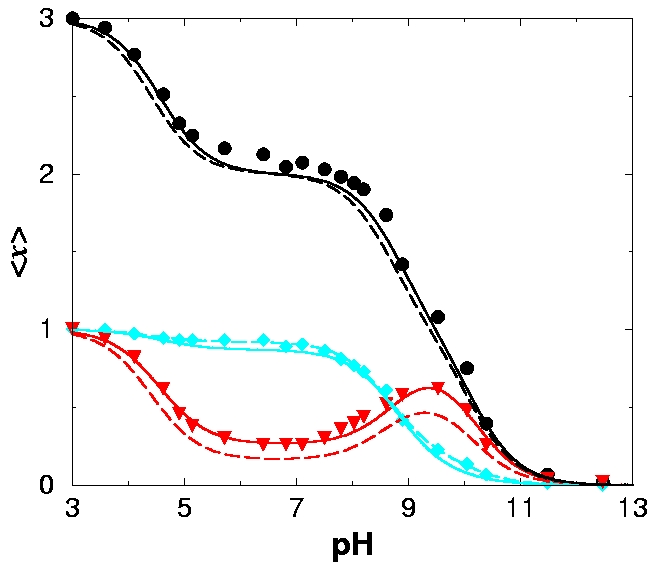
- RIGHT PANEL: Fitting the titration curves of DTPA to the Decoupled Sites Representation (DSR).
The experimental data are represented by
discrete symbols, and the results of the DSR fit are given by lines.
Protonation probabilities are shown on the ordinate.
The colors represent the following protonation probabilities:
red - middle nitrogen,
cyan - terminal nitrogens, and
black - total. Note that the middle nitrogen titration curve (red) is non-monotonic.
Solid lines represent a simultaneous DSR fit to
the two individual titration curves.
For the long-dashed curves, only the data
of the terminal nitrogens
are used (cyan diamonds). Note that fitting the seemingly featureless titration curves
of the terminal nitrogens allows on to reconstruct the non-trivial titration behavior
of the middle nitrogen.
Details can be found in: Alexey Onufriev, D.A. Case and G. M. Ullmann, "A Novel View of pH Titration in Biomolecules", Biochemistry, ("New Concepts") 40, 3413 (2001).