|
| We work at the interface of physics, biology, and computer science; the research is a collective effort of our group members who have background in one or more of these fields.
Most students who graduate from the group become
proficient in cutting edge computing. Our funding comes from the NIH and the NSF.
|
| | Active Research Areas
|
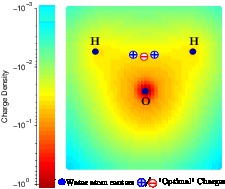 | Better Modeling Accuracy: novel models of water
All biological matter is surrounded by water, but modeling the effects of water accurately and efficiently remains a significant challenge, which we address at various levels. Our most recent efforts combine the power of machine learning with reliability of physics to deliver better models of water. Water -- the simplest most complex molecule -- still holds a number of mysteries, which better water models can help solve.
|
 | Better Medicines, Faster: helping drug discovery efforts through molecular modeling and simulations
Atomistic simulations of biomolecules enable modern drug design efforts; these require fast and accurate algorithms. We harness the power of computer science, including machine learning, to develop better ways to model biomolecules and develop new therapeutics.
|
 | 3D organization of the genome: the basic principles.
The goal is to understand, through a combination
of computation and experiment, the structure-function connections in the genome, which is organized in a complex manner within the cell. Our most recent interests in this direction focus on ageing and cancer. Shown is an artist's vision of the genome organization complexity. Image credit: Eric Standley, VT. |
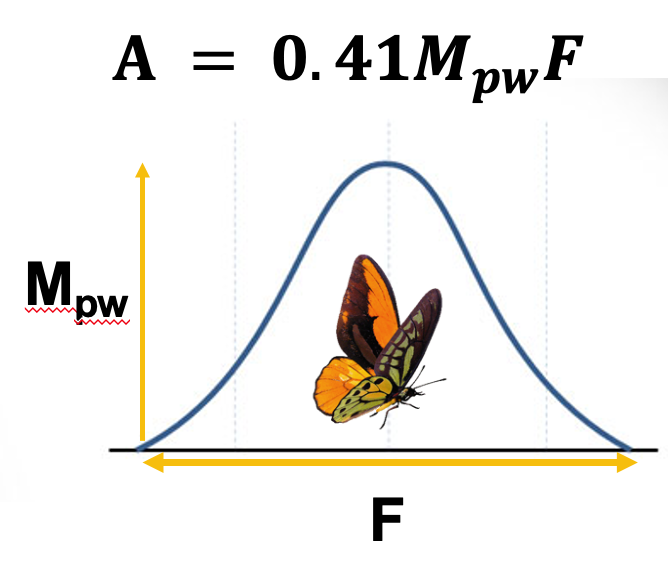 | Mathematical Entomology
Discovering simple mathematical patterns in complex behavior of insects.
|
Other things I worked on in the past
 | DNA compaction and deformation
DNA compaction plays vital role in key cellular processes such as cell differentiation, DNA replication, repair, and transcription.
We aim to uncover the fundamental principles that control the DNA compaction.
|
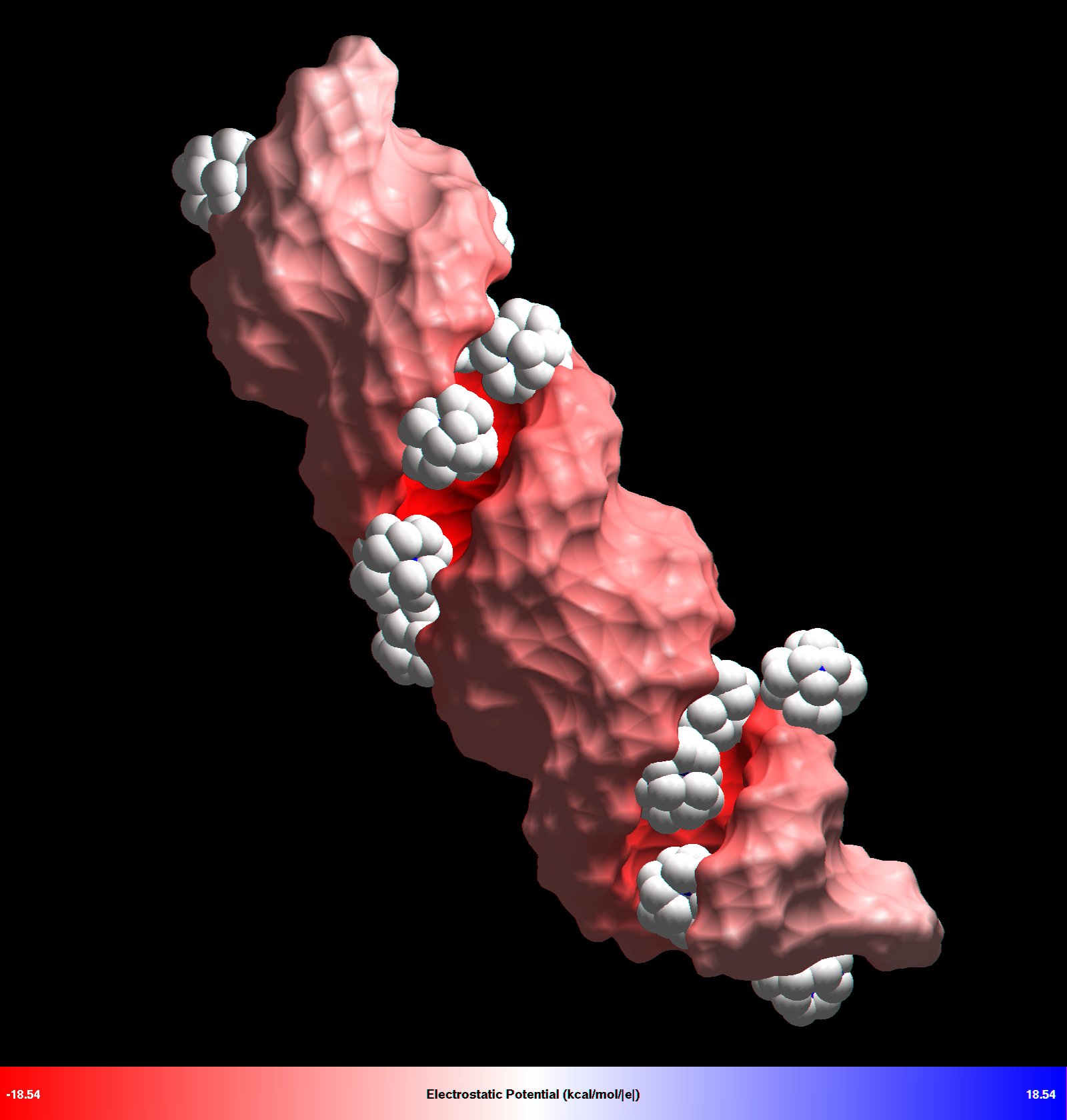 | Nucleic acid condensation induced by ions
Our goal was to develop a quantitative, atomic-level understanding of how multivalent ions, including biologically relevant polyamines, mediate attractive forces between nucleic acid molecules (DNA, RNA) that lead to various nucleic acid condensation phenomena observed in experiments. |
|
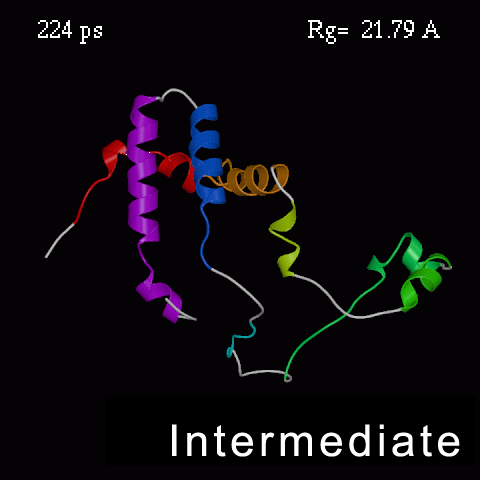 | Protein Folding
One of the largest challenges in modern science is working out how proteins curl up into their complex shapes... |
 | Functional dynamics of biomolecules
Biological function of many biomolecules, such as proteins or DNA, is intricately related to their dynamical properties. Can we understand these relationships using computational methods? |
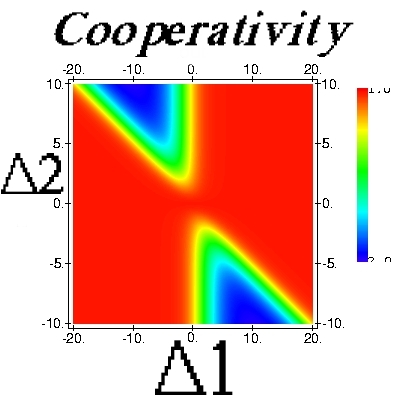 | Theory of Cooperative Ligand Binding
Binding of ligands to macromolecules is one of the most important reactions in biology... |
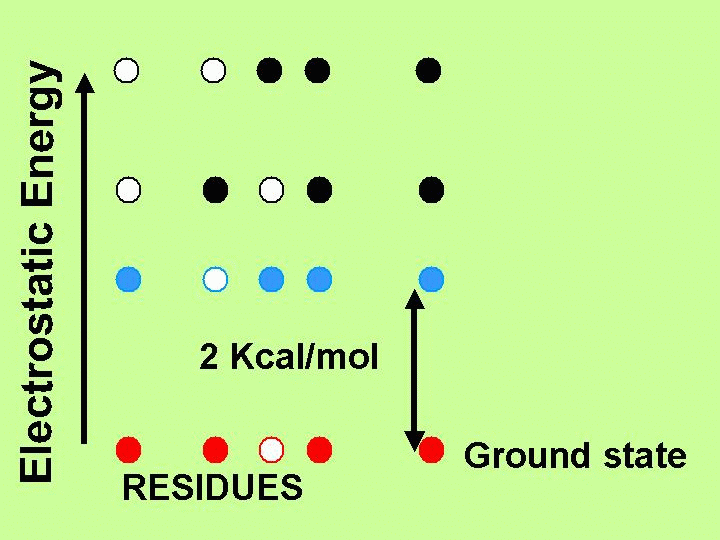 | Proton-pumping mechanism of Bacteriorhodopsin
Bacteriorhodopsin is the smallest autonomous light-harvesting system... |
 | Insights into the RNAi mechanism
Can we use bioinformatics to help us understand how small fragments of RNA interact with the messenger RNA? |
|

