AI & ML for macromolecular modeling:
To this end, we have been developing advanced computational and data-driven methods, specifically leveraging recent breakthroughs in AI & ML, for protein structure prediction. This has resulted in several state-of-the-art methods, ranking among the best methods in multiple rounds of the community-wide CASP (Critical Assessment of Structure Prediction) challenges.
More recently, we have shifted our interest to the modeling of nucleic acids focusing on:
- Deep generative models for sampling RNA conformational ensemble
- Scoring RNA structural models
- RNAbpFlow: Base pair-augmented SE(3)-flow matching for conditional RNA 3D structure generation
- lociPARSE: locality-aware invariant Point Attention-based RNA ScorEr
- QDeep: Protein model quality estimation using deep ResNets
-
RNAbpFlow: Base pair-augmented SE(3)-flow matching for conditional RNA 3D structure generation
Sumit Tarafder, Debswapna Bhattacharya
bioRxiv, 2025
DOI: 10.1101/2025.01.24.634669v1
-
lociPARSE: a locality-aware invariant point attention model for scoring RNA 3D structures
Sumit Tarafder, Debswapna Bhattacharya
Journal of Chemical Information and Modeling, 64 (22), 8655-8664, 2024
DOI: 10.1021/acs.jcim.4c01621
-
QDeep: distance-based protein model quality estimation by residue-level ensemble error classifications using stacked deep residual neural networks
Md Hossain Shuvo, Sutanu Bhattacharya, Debswapna Bhattacharya
Intelligent Systems for Molecular Biology (ISMB), 2020
Bioinformatics, 36(S1): i285-i291, 2020
DOI: 10.1093/bioinformatics/btaa455
LLMs for biomolecular interactions:

We have been developing advanced computational methods powered by pretrained large language models (LLMs) for characterizing biomolecular interactions to enable accurate prediction and validation of biomolecular interactions at scale using embeddings from bio LLMs that can reduce the dependence on the availability of explicit evolutionary information.
Our current interests include:
- Biomolecular binding site prediction
- Single-sequence prediction of biomolecular assemblies
- Accuracy estimation of predicted biomolecular interactions
- EquiPNAS: pLM-informed equivariant graph learning for protein-nucleic acid binding site prediction
- ProRNA3D-single: single-sequence protein-RNA complex structure prediction with biological language models
- EquiPPIS: E(3) equivariant graph neural network for PPI site prediction
-
Single-sequence protein-RNA complex structure prediction by geometric attention-enabled pairing of biological language models
Rahmatullah Roche, Sumit Tarafder, Debswapna Bhattacharya
Cell Systems, In press, 2025
DOI: 10.1101/2024.07.27.605468v1
-
EquiPNAS: improved protein-nucleic acid binding site prediction using protein-language-model-informed equivariant deep graph neural networks
Rahmatullah Roche, Bernard Moussad, Md Hossain Shuvo, Sumit Tarafder, Debswapna Bhattacharya
Nucleic Acids Research, 52 (5), e27-e27, 2024
DOI: 10.1093/nar/gkae039
-
E(3) equivariant graph neural networks for robust and accurate protein-protein interaction site prediction
Rahmatullah Roche, Bernard Moussad, Md Hossain Shuvo, Debswapna Bhattacharya
PLOS Computational Biology, 19(8): e1011435, 2023
DOI: 10.1371/journal.pcbi.1011435
Physics + ML for fine-tuning and refinement:
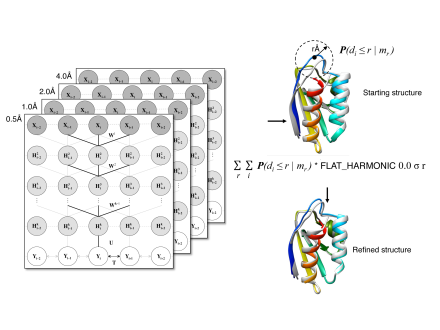
We have developed a number of effective and widely-used refinement methods through optimization of biophysical force fields and/or knowledge-based energy functions coupled with deep learning-derived potentials for improved structure refinement. Free dissemination of the resulting methods and algorithms have attracted promising attention and a broad field-wide impact.
The specific questions we are currently focusing on are:
- Physics-guided ML models for differentiable biomolecular simulation
- ML-guided interatomic potentials for parameterizing atomistic force fields
- 3Drefine: protein structure refinement webserver
- DConStruct: hybridized distance- and contact-based hierarchical structure modeling for folding proteins
-
DeepRefiner: high-accuracy protein structure refinement by deep network calibration
Md Hossain Shuvo, Muhammad Gulfam, Debswapna Bhattacharya
Nucleic Acids Research, Web Server Issue, 49(W1): W147-W152, 2021
DOI: 10.1093/nar/gkab361
-
Hybridized distance- and contact-based hierarchical structure modeling for folding soluble and membrane proteins
Rahmatullah Roche, Sutanu Bhattacharya, Debswapna Bhattacharya
PLOS Computational Biology, 17(2): e1008753, 2021
DOI: 10.1371/journal.pcbi.1008753
-
refineD: improved protein structure refinement using machine learning based restrained relaxation
Debswapna Bhattacharya
Bioinformatics, 35(18): 3320-3328, 2019
DOI: 10.1093/bioinformatics/btz101
Geometric deep learning on molecular graphs:
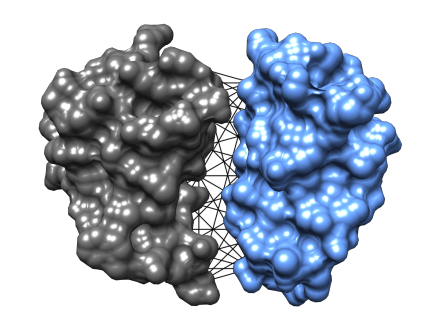
We are developing geometry-aware deep neural networks on molecular graph structured data that preserve symmetries naturally occurring in 3-dimensional space.
We are especially interested in geometric unification of deep representation learning from the perspectives of symmetry and invariance for:
- Principled way to formulate context-specific inductive biases for richer representation of biomolecules
- Explainability and model interpretation
- PIQLE: protein-protein interface quality estimation by deep graph learning
- EquiPPIS: E(3) equivariant graph neural network for PPI site prediction
- EquiPNAS: pLM-informed equivariant graph learning for protein-nucleic acid binding site prediction
-
PIQLE: protein-protein interface quality estimation by deep graph learning of multimeric interaction geometries
Md Hossain Shuvo, Mohimenul Karim, Rahmatullah Roche, Debswapna Bhattacharya
Bioinformatics Advances, 3 (1) vbad070, 2023
DOI: 10.1093/bioadv/vbad070
-
EquiPNAS: improved protein-nucleic acid binding site prediction using protein-language-model-informed equivariant deep graph neural networks
Rahmatullah Roche, Bernard Moussad, Md Hossain Shuvo, Sumit Tarafder, Debswapna Bhattacharya
Nucleic Acids Research, 52 (5), e27-e27, 2024
DOI: 10.1093/nar/gkae039
-
E(3) equivariant graph neural networks for robust and accurate protein-protein interaction site prediction
Rahmatullah Roche, Bernard Moussad, Md Hossain Shuvo, Debswapna Bhattacharya
PLOS Computational Biology, 19(8): e1011435, 2023
DOI: 10.1371/journal.pcbi.1011435
Benchmarking of state of the art:
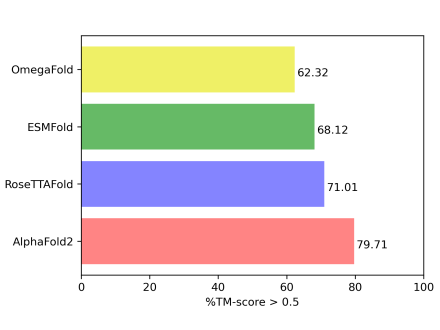
We have been conducting objective and empirical performance benchmarking of the predictive modeling accuracy of the emerging biomolecular modeling methods. The results of our studies provide insights into the current progress while highlighting future areas of improvement.
In our work, we have benchmarked protein and RNA structure prediction methods including:
- Benchmarking protein side-chain packing methods in the post‐AlphaFold era
- Evaluating the landscape of RNA 3D structure modeling with transformer networks
- Assessing the accuracy of AlphaFold2, RoseTTAFold, ESMFold, and OmegaFold in CASP15 targets
- PackBench: benchmarking protein side-chain packing methods in the post‐AlphaFold era
- RNAmark: benchmarking emerging RNA 3D structure prediction methods
- CASP15: benchmarking AlphaFold2, RoseTTAFold, ESMFold, and OmegaFold in CASP15
-
To pack or not to pack: revisiting protein side-chain packing in the post-AlphaFold era
Sriniketh Vangaru, Debswapna Bhattacharya
Briefings in Bioinformatics, 26 (3), bbaf297, 2025
DOI: 10.1093/bib/bbaf297
-
The landscape of RNA 3D structure modeling with transformer networks
Sumit Tarafder, Rahmatullah Roche, Debswapna Bhattacharya
Biology Methods and Protocols, 9 (1), bpae047, 2024
DOI: 10.1093/biomethods/bpae047
-
The transformative power of transformers in protein structure prediction
Bernard Moussad, Rahmatullah Roche, Debswapna Bhattacharya
Proceedings of the National Academy of Sciences of the United States of America, 120 (32) e2303499120, 2023
DOI: 10.1073/pnas.2303499120
© Debswapna Bhattacharya,




